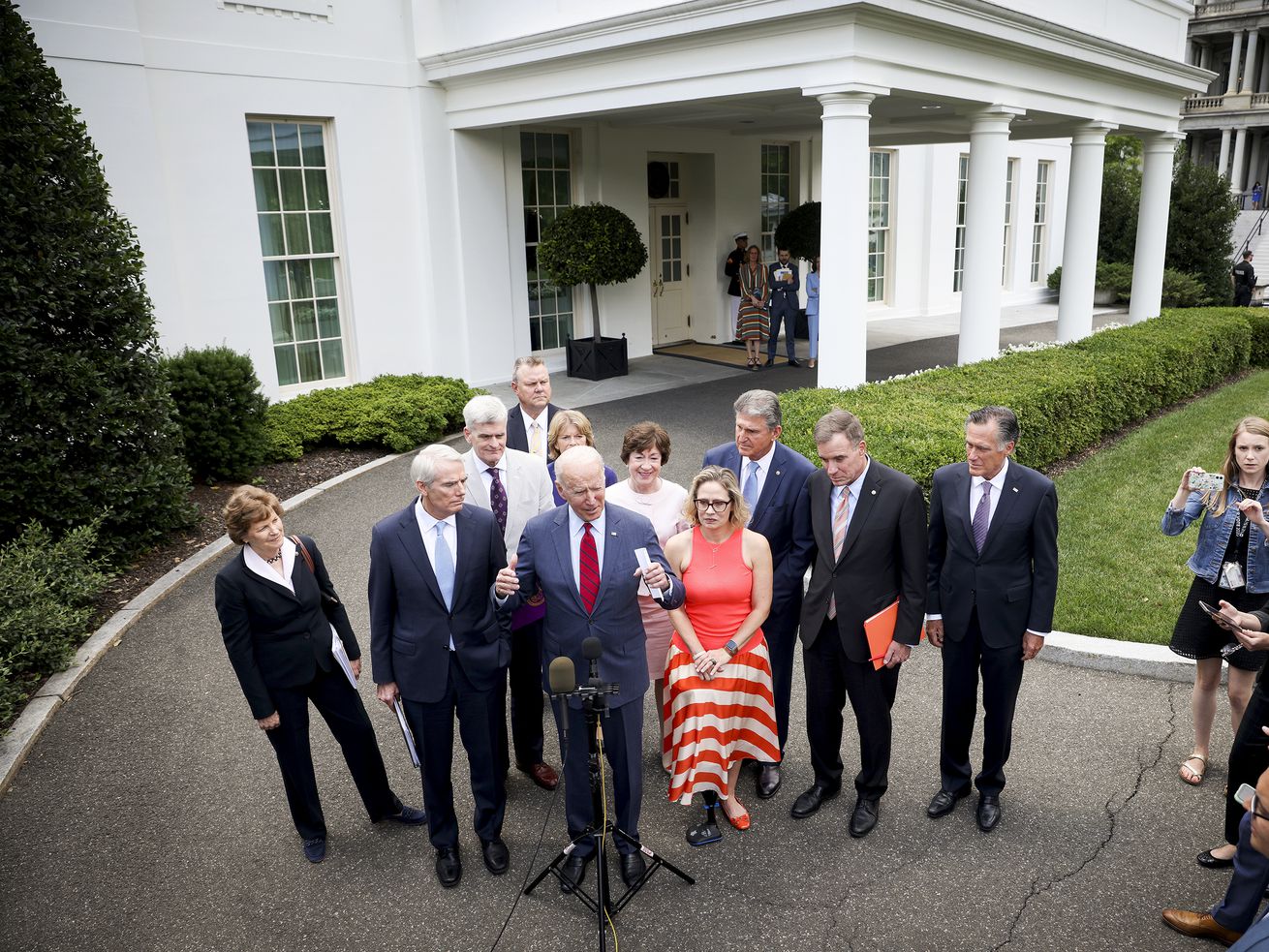
The US lags behind on yet another tool to end the pandemic: viral genetic sequencing.
There’s a reason why a new, more contagious variant of SARS-CoV-2 appeared first in the UK: The country does a lot of viral genetic sequencing. Since the start of the pandemic, researchers in the UK have uploaded 151,859 individual SARS-CoV-2 sequences to GISAID, an international platform for sharing viral genomic data. That’s the highest number of sequences shared by any country in the world.
If a more contagious strain of SARS-CoV-2 first evolved in the United States, scientists likely would not have noticed so quickly. Despite having a larger population than the UK, a sophisticated biomedical research industry, and tens of millions more cases of Covid-19, to date US labs have only uploaded 69,111 sequences, according to GISAID.
“It’s embarrassing, is all I can say,” Diane Griffin, a microbiologist and immunologist at Johns Hopkins, told Vox.
The US has lagged behind on so many aspects of pandemic response — from an initial lack of testing, to the current strained and clumsy rollout of the Covid-19 vaccines. Lack of genetic surveillance is just another. Without it, we’re kept in the dark: Scientists can’t see, clearly or quickly, how and if the virus is mutating in concerning ways. It also leaves us without another useful tool to deploy in contact tracing studies.
And it’s one this country ought to invest in, and get right, scientists say — at least before the next pandemic strikes.
How the US fails on testing viral genomes
Earlier this year, Griffin was on a committee making recommendations for a recent National Academies of Science report on the state of genomic surveillance in the US. Genomic surveillance is used, routinely, around the world to track flu, and to try to predict which flu vaccine strains will be most effective in a given season. Genetic sequencing tools are not a new technology, and the Academies wanted a report to survey how they were being deployed in the pandemic in the US. Genetic sequencing is of particular import when it comes to coronaviruses because they use RNA as their genetic code, and RNA viruses are known to mutate frequently.
The report, when it was published in July, outlined a bleak landscape of SARS-CoV-2 mutation tracking. It’s not just that the US isn’t collecting enough genome samples of the virus. It’s doing so in an unsystematic, patchwork way.
“Current sources of SARS-CoV-2 genome sequence data … are patchy, typically passive, reactive, uncoordinated, and underfunded in the United States,” the report concluded. And the data that did exist? The report found it was “inadequate to answer many of the pressing questions about the evolution and transmission of the virus.”
Early on in the pandemic — way back in March — the UK government invested £20 million ($27 million) to launch the COVID-19 Genomics UK (COG-UK) consortium, which coordinates the collection of this data from public health labs. The consortium also tracks viral genetic samples from health clinics, university research labs, and public health research facilities, to help generate a close-to-real-time snapshot of how the virus is changing in the country.
It’s what allows researchers to generate maps like this one, which shows how the new, more contagious strain of the virus spread geographically in the country over time.
PHE has released the underlying data behind the B.1.1.7 technical report, allowing us to see the spread of the new variant in more detail pic.twitter.com/eSBQzuaIpi
— Theo Sanderson (@theosanderson) January 8, 2021
The rich genetic data, when paired with case reports, also guides researchers to ask and answer crucial questions, such as: Is this new variant more deadly than other ones? Scientists were able to quickly determine the answer is “no.” (That said, a more contagious virus can still end up killing more people than a more virulent one.)
The US Centers for Disease Control and Prevention does have a genetic surveillance program called SPHERES (SARS-CoV-2 Sequencing for Public Health Emergency Response, Epidemiology, and Surveillance), but it’s less well coordinated than the UK effort. Right now labs have to essentially raise their hands and volunteer to contribute. And the funding for their efforts isn’t consistent. That leads to a patchwork of surveillance across the country. “So you might know what’s going on in Boston, or New York City, but have no idea what’s going on in Iowa,” Griffin says.
“In other words,” says Stanford microbiologist David Relman, who also contributed to the National Academies report, “anybody who has the means and interest to engage in genomics is certainly encouraged to do so.” But genomic sequencing, he says, hasn’t been made a “mainstream central pillar of public health efforts.”
What we lose out on when we don’t collect genetic samples of circulating viruses
The National Academies report was published in July. Has the situation gotten much better since? “No,” Griffin says. There has been a little bit of positive movement: Recently, private genomics companies Illumina and Helix have started to help in the detection of new variants in the United States. Even so, James Lu, president of Helix, told MIT Technology Review the US still needs to go from sequencing a few hundred samples a day to around 7,000 per day.
Viral genomics surveillance doesn’t just allow researchers to spot new variants, it helps them learn crucial lessons about how the virus is spreading.
Scientists take advantage of the fact that viruses are constantly making copies of themselves. And every time they make a copy, they may make a little typo in their genetic code. Most of the time, these mutations are meaningless, but they occur at a regular rate. And that makes it possible to make a family tree of the virus. If one viral sample and another have similar typos, researchers can determine they are more closely related.
This can generate key insights.
“In the beginning of the pandemic, we got our hands on some of the first cases that were identified in Connecticut,” says Mary Petrone, a PhD student who works in a molecular biology lab at Yale. Using genomic data, Petrone and her colleagues were able to figure out whether these cases were introduced from abroad, or came from somewhere in the United States. The genetic data revealed that the viruses more closely resembled those circulating on the West Coast than strains from abroad. “It was telling us: there is actually domestic transmission going on,” she says.
Petrone’s lab delivered a key early insight into understanding the virus’s spread in the US. But it wasn’t like the CDC directed them to do so. “Our lab was actually originally set up to do this type of research for mosquito-borne viruses,” she says. “When the pandemic hit we switched over, because there was an urgent public health need to answer some of these questions. So we just happened to really to be set up to do this type of work.”
Setting up more labs to do this work could also help with contact tracing efforts, overall. “For example, if 10 college students test positive,” Julie Segre, a scientist at the National Human Genome Research Institute, writes in an email, “did they come to school already colonized [i.e. infected] or did they transmit the virus while at school.” Genetic evidence can help answer such a question and help prevent future outbreaks.
What needs to happen: coordination, and money
And it’s not necessarily cheap or easy work to do. While the technology that sequences the viral genomes has become relatively inexpensive in recent years (a plug-in USB sequencer will set you back around $1,500), it still takes a lot of skilled lab work to prep samples for analysis. “You definitely don’t need a PhD to be able to do it,” Petrone says. “But you do need to be pretty well trained in molecular biology in the lab. There are a lot of steps where you can contaminate your samples. It can be quite expensive to do.”
Petrone’s lab can do full genome sequencing; that is, they can read every letter of a virus’s genetic code. But not all labs would need to do that to contribute to a surveillance effort. For instance, Petrone’s group is working on a simpler test that can identify the more contagious B117 variant that first was detected in the UK. “That is something you’d be able to run in a clinic,” she says.
But creating a widespread surveillance network for the new variant would require a lot more coordination than what’s currently taking place.
That’s why the US government needs to be more proactive on this, and help set up a nationwide network for genomic data. And that may be coming. According to STAT, the incoming Biden Administration plans to scale up the country’s genomic sequencing efforts as part of a $415 billion emergency Covid-19 spending package it will ask Congress to approve. (Perhaps also auspicious: Biden has selected Eric Lander, a geneticist who co-led the Human Genome Project, to lead the White House Office of Science and Technology Policy, which will be elevated to a Cabinet-level position.)
For a robust genetic surveillance network to be most useful, it needs to be backed up with other rich datasets too. New variants pop up all the time. What matters is whether those variants are linked to worse health outcomes, more reinfections, or faster spread.
“We would ideally have access to good, consistent data about each sample — at the least, geographical location, but more would be better,” Adam Felsenfeld, director of genome sciences at the National Human Genome Research Institute, writes in an email. If possible, too, “one would need details about the medical record of the patients,” he writes, to try to determine if genetic changes in the virus correspond to different disease courses. Again, this would take coordination, as researchers would need informed consent from people to collect this personal data.
A network of viral genome surveillance isn’t just needed for this pandemic, but for future ones too.
“This won’t be the last pandemic,” Griffin says. “If we could get the infrastructure right and get the approach right, then you have things in place you could activate” … for the next time.
Author: Brian Resnick
Read More